. Introduction
An opening of the corolla is an essential stage in higher plants’ life cycle, regulated by a complex set of hormonal and external environmental factors (Larsson et al., 1998; Prudencio et al., 2020). Several pathways have been identified to regulate flowering in the model plant Arabidopsis thaliana (He, 2012), including photoperiod, vernalization, gibberellin, ambient temperature, and age-related pathways.
Polypeptide hormones are important signal transduction molecules in plants and play a crucial role in growth and development (Song et al., 2013). CLE family is a relatively large class of polypeptide signaling molecules in plants (Fletcher et al., 1999; Opsahl-Ferstad et al., 1997; Strabala et al., 2006). The A. thaliana CLE family contains 32 members with conserved CLE motifs and signal peptides (Chu et al., 2006). In vitro, when the small peptide (synthesized from the CLE motif of CLV3) applies, A. thaliana presents a phenotype of gene overexpression, indicating that CLV3 is a secretory peptide (Fiers et al., 2005). CLV3 is mainly expressed in the central region of A. thaliana shoot apical meristem (SAM) as a dynamic apoplast signal and regulates the stem cell development through the CLV3-WUS feedback loop pathway (Brand et al., 2000; Schoof et al., 2000; Shinohara & Matsubayashi, 2013). The root of functional deletion mutant cle40-En becomes shorter and tilted to the left (Hobe et al., 2003). CLE40 can bind to the receptor ACR4, inhibit the expression of WOX5, and maintain the balance between proliferation and differentiation of apical root meristem (RAM) stem cells through CLE40-WOX5 feedback regulatory loop (Stahl et al., 2009). It is also found that CLE41 and CLE44 could further enhance the proliferation of stem cells by promoting the expression of WOX4 (Qiang et al., 2013). CLE8-WOX8 signal transduction pathway is involved in the early development of embryo and endosperm in A. thaliana (Fiume & Fletcher, 2012). AtCLE25 positively regulates water transport from roots to shoots (Takahashi et al., 2018). CLE3 acts as a root-derived long-distance signal that regulates shoots’ systemic acquired resistance (SAR) by interaction with WRKY33 (Ma et al., 2020, 2022).
The CLE family is a relatively large class of polypeptide signaling molecules in plants, and overexpressing CLE19, CLE21, or CLE25 delays the development of rosettes (Strabala et al., 2006). In A. thaliana, AtCLE16, AtCLE17, and AtCLE22 display high promoter activity in sepals and petals at different flower development stages (Jun et al., 2010). However, there is little research on the functions of plant peptides CLE3 in the development of flowers.
A corolla opening is an important process affecting the quality of fruits and market competitiveness in cucumber. We introduced a unique cucumber line, ‘6547’, that possesses extra-long ovaries and shows delayed corolla opening when nutrient supplies are abundant. The extra-long ovary reaches the commercial harvesting stage 2–3 d after flowering while the corolla remains fresh. The obtained marketable cucumber fruits from the two types of the ovary are indistinguishable in length. However, the fruits from extra-long ovaries are straight and have flowers remaining on the tip, which is regarded as more attractive and preferred in the fresh market (Sun et al., 2016). Interestingly, by transcriptome analysis, we have also found that the expression of CsCLE3 (Csa4G627800) negatively correlates with the formation of an extra-long ovary (Sun et al., 2016), suggesting that the gene regulates extra-long ovary formation.
Here, we investigate the function of CsCLE3 and reveal the mechanism of the delayed female corolla opening in cucumbers by conducting transgenic experiments and phenotypic analysis.
. Material and methods
Plant materials and treatments
The cucumber (Cucumis sativus) line ‘6457’ was grown in the greenhouse of Hebei Normal University of Science and Technology under two cultivation treatments described in the previous study (Sun et al., 2016).
The extra-long ovary rate and extra-long ovary plant rate analysis
The extra-long ovary rate and extra-long ovary plant rate were calculated using the following formulas:
Quantitative real-time PCR (qRT-PCR)
Based on color and shape, corolla development can be divided into four stages: green bud, green-yellow bud, yellow bud, and flowering. In the typical ovary, 1 DAL (T1), 3 DAL (T3), 4 DAL (T4), and 5 DAL (T5) indicated the green bud, green-yellow bud, yellow bud, and flowering stages, respectively. In contrast, in the extra-long ovary, these four stages typically corresponded to 1 DAL (EL1), 3 to 5 DAL (EL3, EL4, and EL5), 7 to 8 DAL (EL7 and EL8), and 9 DAL (EL9), respectively. Female corollas at different developmental stages under typical ovary and extra-long ovary conditions were frozen in liquid nitrogen and stored at −80 °C for total RNA extraction (Waryoung, China, http://www.huayueyang.com). First-strand cDNA was synthesized and used as a template for qPCR with an SYBR Premix Taq Mix (Takara) on Applied Biosystems 7500 real-time PCR system. The thermal conditions for real-time PCR were 95 ° for 10 min (denaturation) followed by 40 cycles at 95 ° for 15 s and 60 ° for 1 min.
Alpha-tubulin (TUA, AJ715498) and ubiquitin-like protein (UBI, AF104391) were used as candidates for reference genes (Wan et al., 2009). The expression profiles and stability of cucumber ovary at different physiological developmental stages for two reference genes were analyzed according to the literature (Liu et al., 2020), using three different methods with BestKeeper (Pfaffl et al., 2004), geNorm (Vandesompele et al., 2002), and NormFinder (Andersen et al., 2004) software packages. Primer efficiencies and standard deviations were calculated using qBase software (version 1.3.5) (Hellemans et al., 2007) on a standard curve generated using a five-fold dilution series of one sample over at least six dilution points measured in triplicate. Three biological and three technical repeats were conducted for each gene. Statistical analysis was conducted with two-tailed Student’s t-tests (*P < 0.05, **P < 0.01). Primers are listed in Table S1.
Cucumber transformation
The full-length coding sequence (CDS) of CsCLE3 was inserted into the pCAMBIA1305.1 (Novagen, USA) vector to generate the Pro35S: CsCLE3 overexpression vector. Two target sequences (19 bp) located in the exon were used to construct the CRISPR/Cas9 knockout vector (pKSE402 with GFP fluorescent screening marker, pCBC-DT1T2 as an intermediate vector). The recombinant knockout vector was transformed into Agrobacterium tumefaciens EHA105. The detailed cucumber transformation protocol can be found in the previous study (Hu et al., 2017). Cotyledons were infected by EHA105 (in infection fluid with OD600 = 0.2–0.3) under negative pressure. After three days of co-culture in darkness, explants were transferred to a bud differentiation medium with timentin (200 µg/L) under light/8 h dark at 26 °C for 3–4 weeks (Ding et al., 2015). Then the buds with the GFP marker were selected, excised from an explant, and transferred to a rooting medium (Ding et al., 2015). The homozygous T1 mutants without vector (GFP-free) were identified from T0 transgenic lines for further phenotype observation and data statistics. Primers are listed in Table S1.
. Results
Expression patterns of CsCLE3
To check the stability of the candidate reference genes TUA and UBI in a cucumber ovary at physiological development stages, we used RT-qPCR. We observed the expression of the two reference genes at the T1, T3, and EL7. The results showed that the Ct values of the two candidates were between 15 and 21 at T1, T3, and EL7 (Figure S1A). The geNorm, NormFinder, and BestKeeper stability analysis showed that TUA and UBI are stable reference genes (Figure S1B-E).
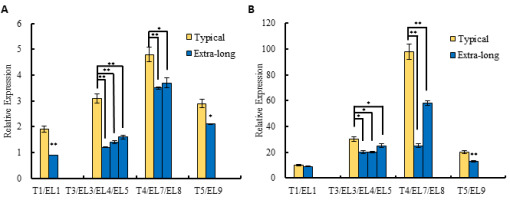
In this study, the relative expression levels of CsCLE3 in the female flowers at different developmental stages under the typical ovary and the extra-long ovary conditions of ‘6457’ were measured by qRT-PCR. We found that CsCLE3 had similar expression patterns both under the typical ovary and the extra-long ovary conditions based on the data of transcriptome analysis and qRT-PCR (Figure 1). The expression of CsCLE3 increased significantly at the stage of yellow bud (N7/S7/S8) and then decreased significantly. However, the expression level of CsCLE3 in the extra-long ovary was significantly lower than that of the typical ovary during the female corolla opening.
Figure 1
Relative expression of CsCLE3 at different stages of corolla opening in ‘6457’. (A) Transcriptome analysis (Sun et al., 2016). (B) The qRT-PCR analysis. Two-tailed Student’s t-tests were used (*P < 0.05, ** P < 0.01). The four stages of corolla development in the typical ovary (T) and extra-long ovary (EL) at which samples were collected for analysis: green bud, green-yellow bud, yellow bud, and flowering. In the typical ovary, 1 DAL (T1), 3 DAL (T3), 4 DAL (T4), and 5 DAL (T5) indicated the green bud, green-yellow bud, yellow bud, and flowering stages, respectively. In contrast, in the extra-long ovary, these four stages typically corresponded to 1 DAL (EL1), 3 to 5 DAL (EL3, EL4, and EL5), 7 to 8 DAL (EL7 and EL8), and 9 DAL (EL9), respectively.
CsCLE3 Negatively regulates extra-long ovary formation in cucumber
To identify the biological functions of CsCLE3 in cucumber, the Pro35S: CsCLE3 overexpression vector contained the full-length CDS of CsCLE3 driven by the CaMV 35S promoter, was transformed into ‘6457’. The representative line Cscle3-oe-11 was used for further characterization. Our data showed that the overexpression of CsCLE3 inhibited the formation of extra-long ovaries. Both the average extra-long ovary rate and the extra-long ovary plant rate were 0% (Figure 2B, Table 1). The overexpressed line Cscle3-oe-11 mostly entered anthesis at 5 d after labeling (DAL; labeling was done when an ovary became visible) (Figure 2C).
Figure 2
Morphological characterization of the transformation lines. (A) Mutation form of T1 transgenic Cscle3-cr-2 line by CRISPR/Cas9 system. (B) Ovary and corolla phenotype at the day of corolla open in ‘6457’, Cscle3-oe-11, and Cscle3-cr-2. Quantification of the days to anthesis (C), ovary length at the day of corolla open (D), and development rates of ovary length in ‘6457’, Cscle3-oe-11, and Cscle3-cr-2 (E). Scale bar = 1 cm.
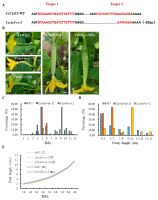
Table 1
Extra-long ovary rate and extra-long ovary plant rate for different lines.
| Line | Extra-long ovary rate (%) | Extra-long ovary plant rate (%) |
|---|---|---|
| 6457 | 29.38 | 24.05 |
| Cscle3-oeor-11 | 0 | 0 |
| Cscle3-cr-2 | 66.67 | 100.00 |
To confirm the function of CsCLE3 in cucumber, we constructed CsCLE3-knockout lines in ‘6457’ by the CRISPR-Cas9 gene-editing system. The two targets are located in the exon. One loss-of-function mutant line (Cscle3-cr-2) was selected for further characterization (Figure 2A). The knockout of CsCLE3 in ‘6457’ could have enhanced the extra-long ovary phenotype. The average extra-long ovary rate was 66.67%, and the average extra-long ovary plant rate was 100% (Figure 2B, Table 1). According to the quantification of the days to blooming, we found that the extra-long ovary of Cscle3-cr-2 mostly entered anthesis at 8–10 DAL, which was 4-d to 5-d delayed in comparison to the typical ovary (Figure 2C).
For the typical (Cscle3-oe-11) and the extra-long ovary (Cscle3-cr-2) of ‘6457’, the average ovary length on the day of the corolla opening was 5.4 cm and 11.8 cm, respectively (Figure 2D). However, among the three lines, the ovary growth rates and length at the marketable stage of the two types of ovaries correspond to each other (Figure 2E).
. Discussion
A corolla opening is an evolutionary breakthrough in the reproduction of higher plants (van Doorn & Kamdee, 2014). Nevertheless, the molecular mechanism of a corolla opening remains largely unknown, mainly due to the lack of mutants or lines that display appreciable changes during a corolla opening. In cucumber, the time of the female flower anthesis and corolla durability is a vital appearance quality trait that requires extensive investigation. For most cucumber cultivars, this process takes 4d to 5d and is insensitive to nutrient conditions (Sun et al., 2016). In cucumber inbred line ‘6547’, the corolla opening of flowers with the extra-long ovary was 4–5 days delayed compared to flowers with the typical ovary. However, the female flower anthesis’s main genes and molecular mechanism remain unknown. In this study, we found through transgenic experiments that CsCLE3 delays the female flower anthesis in cucumbers. These findings deepen our knowledge of the female flower anthesis and provide new insight into the basis of molecular cucumber reproduction, producing fruits with flowers remaining on the tip.


